| 产品货号 | 载体名称 | 出品公司 | 载体用途 | 产品价格 | 出货周期 |
|---|---|---|---|---|---|
| HG-VJC0483 | pGADT7 / pGADT7-AD | Clontech |
酵母双杂交系统
|
询价 | 现货 |
质粒类型:酵母双杂交系统
启动子:ADH1
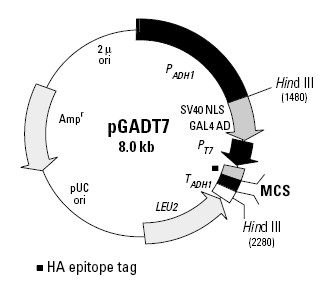
载体大小:8.0kb
载体标签:HA
载体抗性:Ampicillin (氨苄青霉素)
The pGADT7 vector expresses proteins fused to amino acids 768–881 of the GAL4 activation domain(AD). In yeast, fusion proteins are expressed at high levels from the constitutive ADH1 promoter(PADH1); transcription is terminated at the ADH1 transcription termination signal (TADH1). The fusion protein is targeted to the yeast nucleus by the SV40 nuclear localization sequences that have been added to the activation domain sequence. pGADT7 also contains the T7 promoter, an HA epitope tag, and a MCS. pGADT7 replicates autonomously in both E. coli and S. cerevisiae from the pUC and 2 u ori, respectively. The vector carries Ampr for selection in E. coli and the LEU2 nutritional marker for selection in yeast.
pGADT7 is the AD Vector included with MATCHMAKER Two-Hybrid System 3. The MCS of pGADT7 has unique restriction sites in frame with the 3'-end of the GAL4 AD for constructing a fusion protein with either a protein of interest or a fusion protein library. The bait protein is also expressed as a fusion to a hemagglutinin (HA) epitope tag. HA-tagged proteins can be identified with antibodies raised to this common epitope, eliminating the need to generate specific antibodies to new proteins. The T7 promoter is used for in vitro transcription and translation of the epitope tagged fusion protein and also provides a binding site for sequencing using the T7 Sequencing Primer. Note that the AD is not expressed during the in vitro transcription and translation reactions.
The Nco I and Pst I sites may be used to shuttle inserts from pGADT7 into pGBKT7, the MATCHMAKER Two-Hybrid System 3 DNA-BD Vector. The MCS in pGADT7 is compatible with those in pMyc-CMV and pHA-CMV, CLONTECH's epitope tagged mammalian expression vector set (#K6003-1). As a result, the target gene can be shuttled into these vectors in order to confirm protein interactions in vivo.
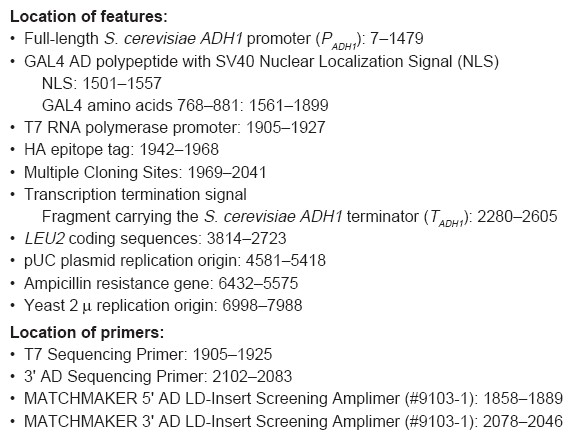
Location of features:
Full-length S. cerevisiae ADH1 promoter (PADH1): 7–1479
GAL4 AD polypeptide with SV40 Nuclear Localization Signal (NLS)
NLS: 1501–1557
GAL4 amino acids 768–881: 1561–1899
T7 RNA polymerase promoter: 1905–1927
HA epitope tag: 1942–1968
Multiple Cloning Sites: 1969–2041
Transcription termination signal
Fragment carrying the S. cerevisiae ADH1 terminator (TADH1): 2280–2605
LEU2 coding sequences: 3814–2723
pUC plasmid replication origin: 4581–5418
Ampicillin resistance gene: 6432–5575
Yeast 2 m replication origin: 6998–7988
Location of primers:
T7 Sequencing Primer: 1905–1925
3' AD Sequencing Primer: 2102–2083
MATCHMAKER 5' AD LD-Insert Screening Amplimer (#9103-1): 1858–1889
MATCHMAKER 3' AD LD-Insert Screening Amplimer (#9103-1): 2078–2046
Propagation in E. coli:
Suitable host strains: DH5a, DH10 & other general purpose strains
Selectable marker: plasmid confers resistance to ampicillin (100 mg/ml) to E. coli hosts
E. coli replication origin: pUC
Copy number: ~500
Plasmid incompatibility group: pMB1/Col E1
Propagation in S. cerevisiae:
Suitable host strains: Y187(a), Y190(a), SFY526(a), CG1945(a), HF7c(a), or AH109(a)
Selectable marker: LEU2
S. cerevisiae origin: 2 u